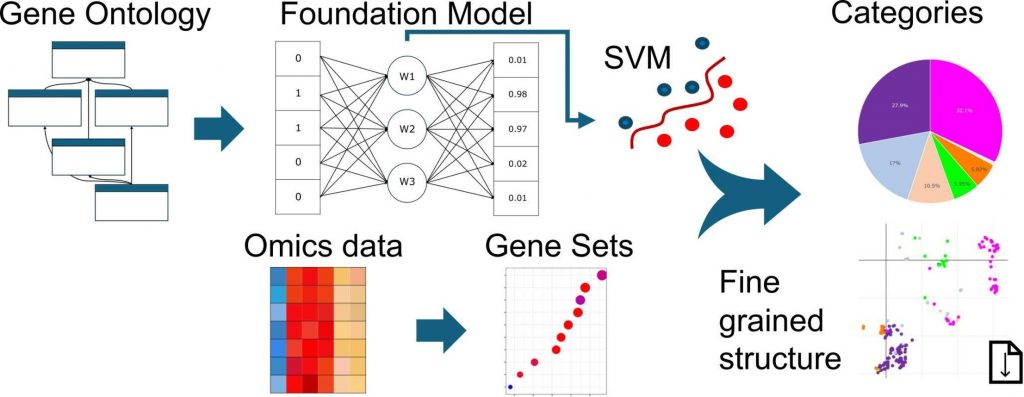
Gene Ontology (GO) is the most commonly used scientific knowledge base, providing functional annotations for gene sets. The ever-growing number of high-throughput biological experiments identifying differentially expressed genes (DEG) statistically enriched in those gene sets often results in long and redundant lists of GO terms. To facilitate the analysis and subsequently visualize trends within those lists of gene sets, we propose a database derived from a natural language processing-based machine learning algorithm that classifies GO terms into predefined higher-order groups. To support viral research, we predefined 18 such groups of higher-order annotation and provided a database of categorized GO terms tailored to characterize the behavior of biological systems in the context of viral infection.